Researchers at the University of Michigan have created a statistical technique that can be applied to various purposes, including mapping your ancestry, modeling the dissemination of diseases, and analyzing how species distribute across geographic landscapes.
One application of this technique is to provide a more comprehensive representation of human lineage, according to Gideon Bradburd, a professor of ecology and evolutionary biology at U-M. For instance, when you submit your DNA for a personalized ancestry analysis, the report returned gives merely a narrow perspective of your family tree positioned at a specific moment in time and space.

Such genetic analyses reflect the portion of a person’s genome that is inherited from ancestors who lived in a specific region during a particular moment in history. For example, if your ancestry report indicates that you are 50% Irish, it suggests that you have numerous second through fourth cousins presently residing in Ireland, explains Bradburd. However, your family tree is ultimately much more dynamic than this singular snapshot.
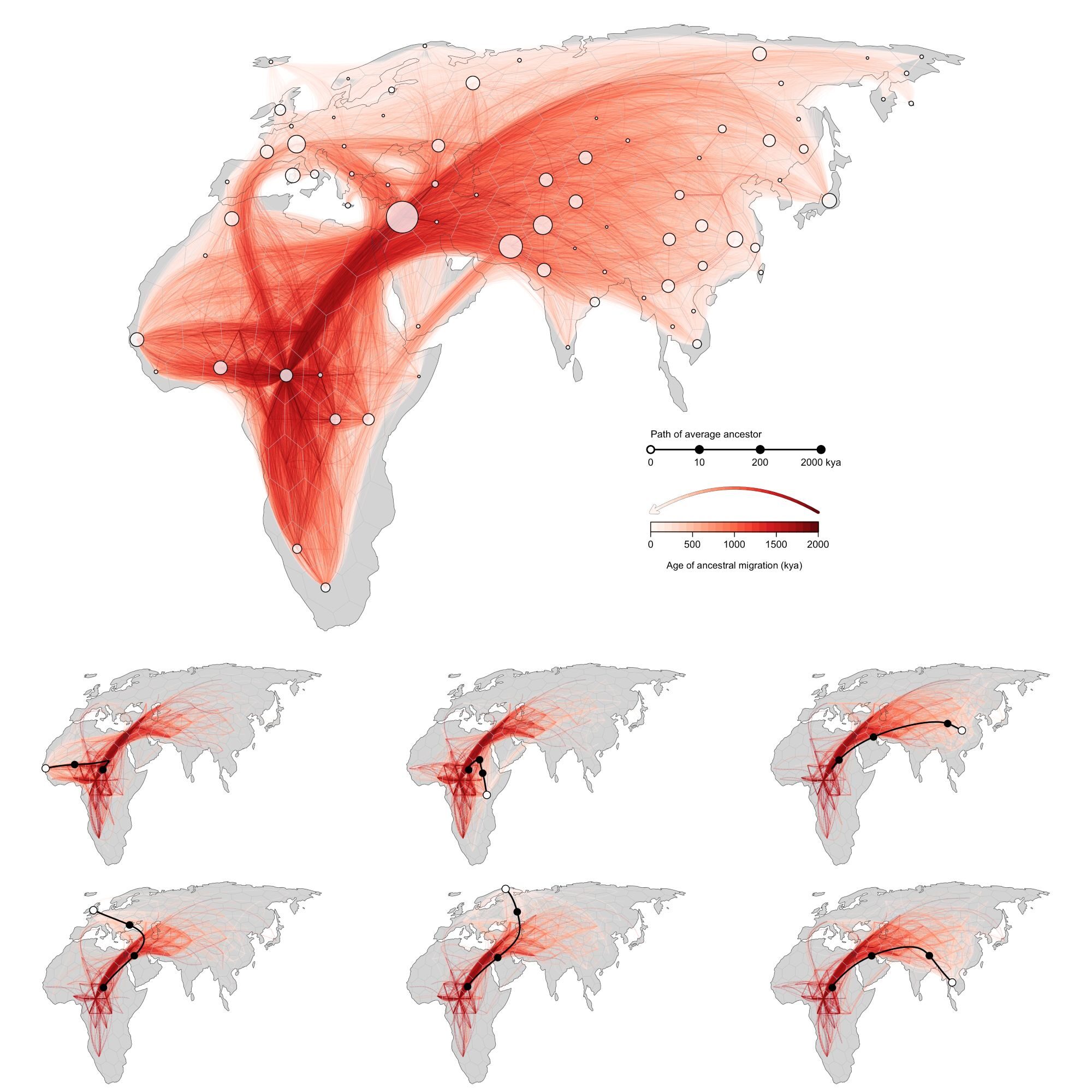
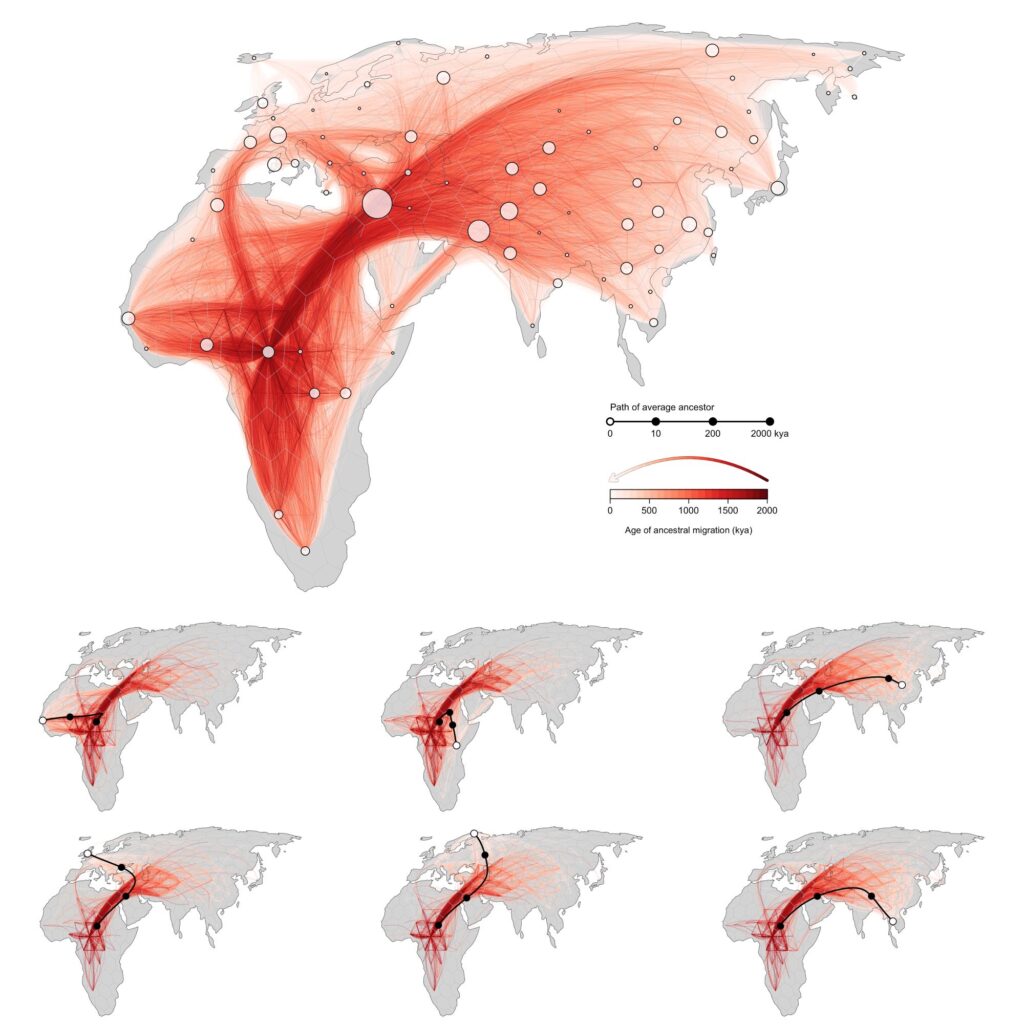
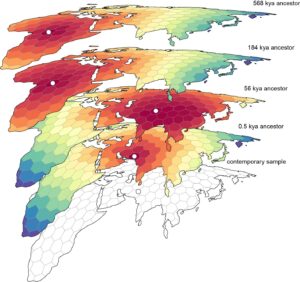
The statistical approach devised by Bradburd alongside fellow researchers at U-M, Michael Grundler and Jonathan Terhorst, can present individuals with a “movie” depiction of their ancestry, illustrating the origins of their forebears and their movements throughout the globe. The method leverages modern genetic sequence data, estimates all locations of an individual’s genetic forebears, identifies the average location of those individuals based on hypotheses about migration patterns, and traces it back over centuries.

The methodology developed by the researchers extends beyond simply tracing human ancestry. It can also be applied to monitor the emergence of viruses, the divergence of animal populations, and other genealogical tracking. Their findings are published in the journal Science.
“There are ways in which consumer ancestry reports are captivating, and certainly it’s enlightening to uncover your history, especially for individuals who’ve been adopted, orphaned, or are disconnected from their families,” Bradburd remarked. “Yet, these reports can also pose substantial issues. They reinforce perceptions of biological essentialism related to race, as they present categories like Irish as if they are fixed ideals that remain unchanged over time.”

However, the researchers are aware that this is not accurate. A field established by Nobel Prize-winning geneticist Svante Pääbo developed the technology to genotype ancient DNA, enabling scientists to trace waves of human populations as they migrated across the globe—especially in Eurasia, where a majority of this type of genetic sequencing has been conducted, according to Bradburd. This innovation allows scientists to see how human groups enter and exit geographic regions over time.
“Since the genetic composition of a location shifts significantly throughout history, we realize that it is inconsequential to assert, ‘This is what it means to be Irish,’” Bradburd explained. “It’s not merely that being ‘genetically Irish’ holds no substantial meaning; it implies that you embody everything.”
Bradburd highlights an illustrative thought experiment in human genetics: envision two biological parents and four biological grandparents. This doubles every generation, and it only takes a relatively small number of generations before there are more individuals in that direct lineage than there have ever been humans living on Earth.
This also signifies that you do not need to reach very far back in time to find that numerous individuals share a multitude of ancestors.
“As our genealogies expand so rapidly, they must also condense in the same way that you and I must share countless relatives at various points in history, and this is true for every individual inhabiting the planet,” Bradburd commented. “We are all remarkably closely interconnected.”
Bradburd stated that while the ancestry reports are precise, they are particularly relevant when linked to specific historical periods.
“While the ancestry analyses aren’t incorrect, they omit a crucial aspect: the ‘when’ of your Irish heritage,” stated Bradburd. “Since we understand the modern human lineage originated in Africa, it is equally valid to state that you possess 100% African lineage from a longer temporal perspective.”
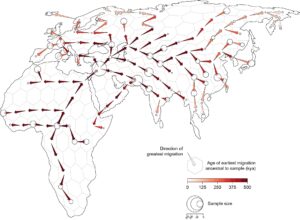
The analytical technique, known as Gaia (geographic ancestry inference algorithm), begins with a straightforward assumption regarding individual movement: predominantly, individuals migrate locally. This technique combines that presumption with the locations of current individuals and a genetic framework that connects them, referred to as the ancestral recombination graph.
Armed with these two elements and the basic model of individual movement, the researchers can determine the “most parsimonious locations of ancestors,” affirms Bradburd. Subsequently, the researchers can trace that information backward through history.
Bradburd’s research addresses a call from the National Academy of Sciences, urging scholars in human population genetics to transition away from racially-defined labels. Although the sociological implications of race are undeniable, racial categories do not effectively predict genetic diversity, according to him.
Due to the disconnection between race and genetics, racial labels can frequently be inaccurate: two individuals may share the same designation, yet be significantly more or less related. Furthermore, since the genetic composition of a particular geographical area can fluctuate considerably over time, geographical and national labels can also be misleading, adds Bradburd.
“Claiming you are ‘genetically Irish’ implies that ‘Irish’ has always held the same meaning, but genetically we recognize that this is false, and anyone deemed Irish—meaning they have inherited portions of their genome from individuals who resided in Ireland—is solely Irish within a specific temporal context,” he remarked. “The fact that these racial labels overlook both of these critical points results in a significant loss of detail and presents a genuine risk of science being exploited for political agendas.”
The approach developed by Bradburd’s team can be utilized for areas beyond human genetics. This method can be employed to analyze the genetic distribution of other organisms in their research. Scholars can also apply this technique to examine the migration patterns of various species—human and otherwise, Bradburd mentions.
For instance, researchers have succeeded in analyzing the genetic similarities of populations between different areas to deduce if they are more or less closely linked via migration or are considerably isolated from one another. However, this tool enables researchers to establish a timeline for when these migrations took place.
Moreover, this instrument is applicable not only for tracing human genetics but could also assist in determining when a particular disease may have originated from a certain area of the globe, for instance. The U-M team collaborates with researchers in Australia to study how mosquitoes populated islands in the South Pacific and with researchers in Michigan and Ohio to explore the history and distribution of the Massasauga rattlesnake.
“I find it exciting that this methodology can be employed to identify dispersal patterns over time and among specific locales,” Bradburd expressed. “Concepts of ancestry need not be static. Rather, they should be viewed as dynamic, captivating, and interpretable more akin to a film than a single image.”